3.5. Descriptor "se_atten"
3.5.1. DPA-1: Pretraining of Attention-based Deep Potential Model for Molecular Simulation
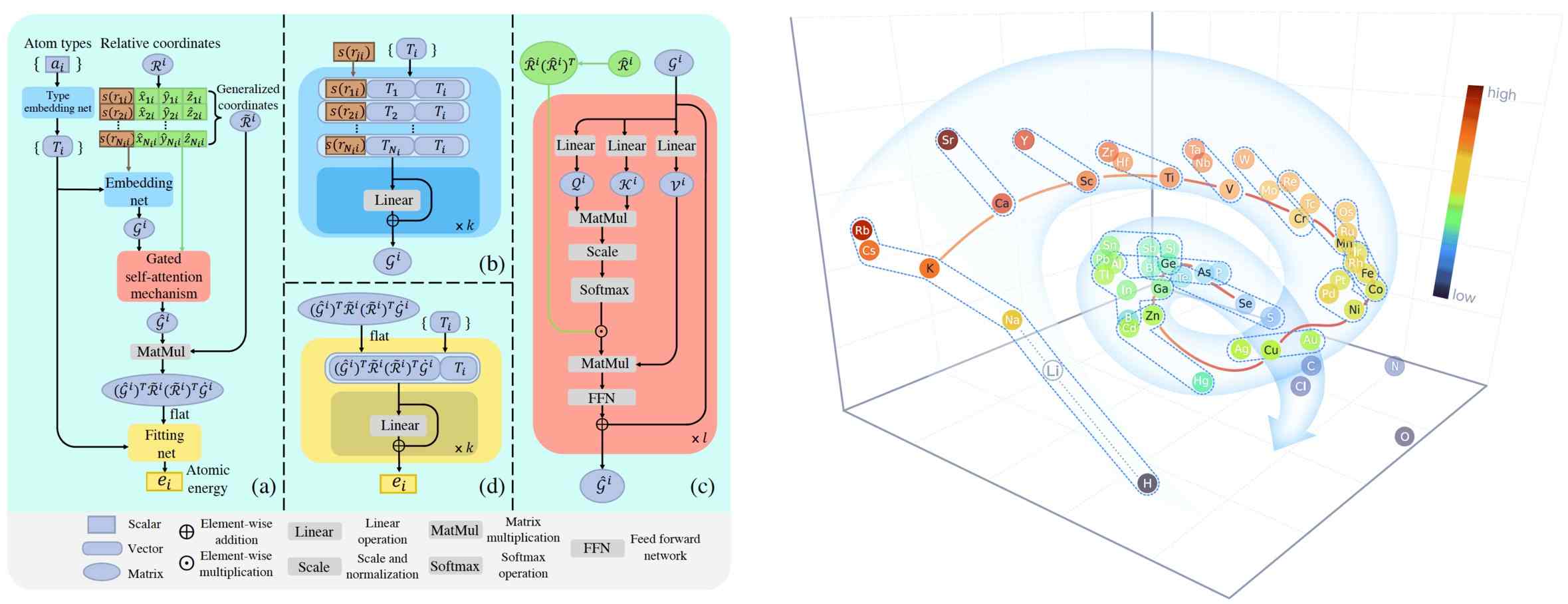
Here we propose DPA-1, a Deep Potential model with a novel attention mechanism, which is highly effective for representing the conformation and chemical spaces of atomic systems and learning the PES.
See this paper for more information. DPA-1 is implemented as a new descriptor "se_atten" for model training, which can be used after simply editing the input.json.
3.5.2. Theory
Attention-based descriptor \(\mathcal{D}^i \in \mathbb{R}^{M \times M_{<}}\), which is proposed in pretrainable DPA-1 model, is given by
where \(\hat{\mathcal{G}}^i\) represents the embedding matrix \(\mathcal{G}^i\) after additional self-attention mechanism and \(\mathcal{R}^i\) is defined by the full case in the se_e2_a. Note that we obtain \(\mathcal{G}^i\) using the type embedding method by default in this descriptor.
To perform the self-attention mechanism, the queries \(\mathcal{Q}^{i,l} \in \mathbb{R}^{N_c\times d_k}\), keys \(\mathcal{K}^{i,l} \in \mathbb{R}^{N_c\times d_k}\), and values \(\mathcal{V}^{i,l} \in \mathbb{R}^{N_c\times d_v}\) are first obtained:
where \(Q_{l}\), \(K_{l}\), \(V_{l}\) represent three trainable linear transformations that output the queries and keys of dimension \(d_k\) and values of dimension \(d_v\), and \(l\) is the index of the attention layer. The input embedding matrix to the attention layers, denoted by \(\mathcal{G}^{i,0}\), is chosen as the two-body embedding matrix.
Then the scaled dot-product attention method is adopted:
where \(\varphi\left(\mathcal{Q}^{i,l}, \mathcal{K}^{i,l},\mathcal{R}^{i,l}\right) \in \mathbb{R}^{N_c\times N_c}\) is attention weights. In the original attention method, one typically has \(\varphi\left(\mathcal{Q}^{i,l}, \mathcal{K}^{i,l}\right)=\mathrm{softmax}\left(\frac{\mathcal{Q}^{i,l} (\mathcal{K}^{i,l})^{T}}{\sqrt{d_{k}}}\right)\), with \(\sqrt{d_{k}}\) being the normalization temperature. This is slightly modified to incorporate the angular information:
where \(\hat{\mathcal{R}}^{i} \in \mathbb{R}^{N_c\times 3}\) denotes normalized relative coordinates , \(\hat{\mathcal{R}}^{i}_{j} = \frac{\boldsymbol{r}_{ij}}{\lVert \boldsymbol{r}_{ij} \lVert}\) and \(\odot\) means element-wise multiplication.
Then layer normalization is added in a residual way to finally obtain the self-attention local embedding matrix \(\hat{\mathcal{G}}^{i} = \mathcal{G}^{i,L_a}\) after \(L_a\) attention layers:1
3.5.3. Introduction to new features of DPA-1
Next, we will list the detailed settings in input.json and the data format, especially for large systems with dozens of elements. An example of DPA-1 input can be found here.
3.5.3.1. Descriptor "se_atten"
The notation of se_atten is short for the smooth edition of Deep Potential with an attention mechanism. This descriptor was described in detail in the DPA-1 paper and the images above.
In this example, we will train a DPA-1 model for a water system. A complete training input script of this example can be found in the directory:
$deepmd_source_dir/examples/water/se_atten/input.json
With the training input script, data are also provided in the example directory. One may train the model with the DeePMD-kit from the directory.
An example of the DPA-1 descriptor is provided as follows
"descriptor" :{
"type": "se_atten",
"rcut_smth": 0.50,
"rcut": 6.00,
"sel": 120,
"neuron": [25, 50, 100],
"axis_neuron": 16,
"resnet_dt": false,
"attn": 128,
"attn_layer": 2,
"attn_mask": false,
"attn_dotr": true,
"seed": 1
}
The type of the descriptor is set to
"se_atten", which will use DPA-1 structures.rcut is the cut-off radius for neighbor searching, and the rcut_smth gives where the smoothing starts.
sel gives the maximum possible number of neighbors in the cut-off radius. It is an int. Note that this number highly affects the efficiency of training, which we usually use less than 200. (We use 120 for training 56 elements in OC2M dataset)
The neuron specifies the size of the embedding net. From left to right the members denote the sizes of each hidden layer from the input end to the output end, respectively. If the outer layer is twice the size of the inner layer, then the inner layer is copied and concatenated, then a ResNet architecture is built between them.
The axis_neuron specifies the size of the submatrix of the embedding matrix, the axis matrix as explained in the DeepPot-SE paper
If the option resnet_dt is set to
true, then a timestep is used in the ResNet.seed gives the random seed that is used to generate random numbers when initializing the model parameters.
attn sets the length of a hidden vector during scale-dot attention computation.
attn_layer sets the number of layers in attention mechanism.
attn_mask determines whether to mask the diagonal in the attention weights and False is recommended.
attn_dotr determines whether to dot the relative coordinates on the attention weights as a gated scheme, True is recommended.
3.5.3.2. Descriptor "se_atten_v2"
We highly recommend using the version 2.0 of the attention-based descriptor "se_atten_v2", which is inherited from "se_atten" but with the following parameter modifications:
"stripped_type_embedding": true,
"smooth_type_embdding": true,
"set_davg_zero": false
Practical evidence demonstrates that "se_atten_v2" offers better and more stable performance compared to "se_atten".
3.5.3.3. Fitting "ener"
DPA-1 only supports "ener" fitting type, and you can refer here for detailed information.
3.5.3.4. Type embedding
DPA-1 only supports models with type embeddings. And the default setting is as follows:
"type_embedding":{
"neuron": [8],
"resnet_dt": false,
"seed": 1
}
You can add these settings in input.json if you want to change the default ones, see here for detailed information.
3.5.3.5. Type map
For training large systems, especially those with dozens of elements, the type determines the element index of training data:
"type_map": [
"Mg",
"Al",
"Cu"
]
which should include all the elements in the dataset you want to train on.
3.5.4. Data format
DPA-1 supports the standard data format, which is detailed in data-conv.md and system.md. Note that in this format, only those frames with the same fingerprint (i.e. the number of atoms of different elements) can be put together as a unified system. This may lead to sparse frame numbers in those rare systems.
An ideal way is to put systems with the same total number of atoms together, which is the way we trained DPA-1 on OC2M. This system format, which is called mixed_type, is proper to put frame-sparse systems together and is slightly different from the standard one. Take an example, a mixed_type may contain the following files:
type.raw
type_map.raw
set.*/box.npy
set.*/coord.npy
set.*/energy.npy
set.*/force.npy
set.*/real_atom_types.npy
This system contains Nframes frames with the same atom number Natoms, the total number of element types contained in all frames is Ntypes. Most files are the same as those in standard formats, here we only list the distinct ones:
ID | Property | File | Required/Optional | Shape | Description |
|---|---|---|---|---|---|
/ | Atom type indexes (place holder) | type.raw | Required | Natoms | All zeros to fake the type input |
type_map | Atom type names | type_map.raw | Required | Ntypes | Atom names that map to atom type contained in all the frames, which is unnecessart to be contained in the periodic table |
type | Atom type indexes of each frame | real_atom_types.npy | Required | Nframes * Natoms | Integers that describe atom types in each frame, corresponding to indexes in type_map. |
With these edited files, one can put together frames with the same Natoms, instead of the same formula (like H2O). Note that this mixed_type format only supports se_atten descriptor.
To put frames with different Natoms into the same system, one can pad systems by adding virtual atoms whose type is -1. Virtual atoms do not contribute to any fitting property, so the atomic property of virtual atoms (e.g. forces) should be given zero.
The API to generate or transfer to mixed_type format is available on dpdata for a more convenient experience.
3.5.5. Training example
Here we upload the AlMgCu example shown in the paper, you can download it here: Baidu disk; Google disk.
- 1
This section is built upon Jinzhe Zeng, Duo Zhang, Denghui Lu, Pinghui Mo, Zeyu Li, Yixiao Chen, Marián Rynik, Li’ang Huang, Ziyao Li, Shaochen Shi, Yingze Wang, Haotian Ye, Ping Tuo, Jiabin Yang, Ye Ding, Yifan Li, Davide Tisi, Qiyu Zeng, Han Bao, Yu Xia, Jiameng Huang, Koki Muraoka, Yibo Wang, Junhan Chang, Fengbo Yuan, Sigbjørn Løland Bore, Chun Cai, Yinnian Lin, Bo Wang, Jiayan Xu, Jia-Xin Zhu, Chenxing Luo, Yuzhi Zhang, Rhys E. A. Goodall, Wenshuo Liang, Anurag Kumar Singh, Sikai Yao, Jingchao Zhang, Renata Wentzcovitch, Jiequn Han, Jie Liu, Weile Jia, Darrin M. York, Weinan E, Roberto Car, Linfeng Zhang, Han Wang, J. Chem. Phys. 159, 054801 (2023) licensed under a Creative Commons Attribution (CC BY) license.